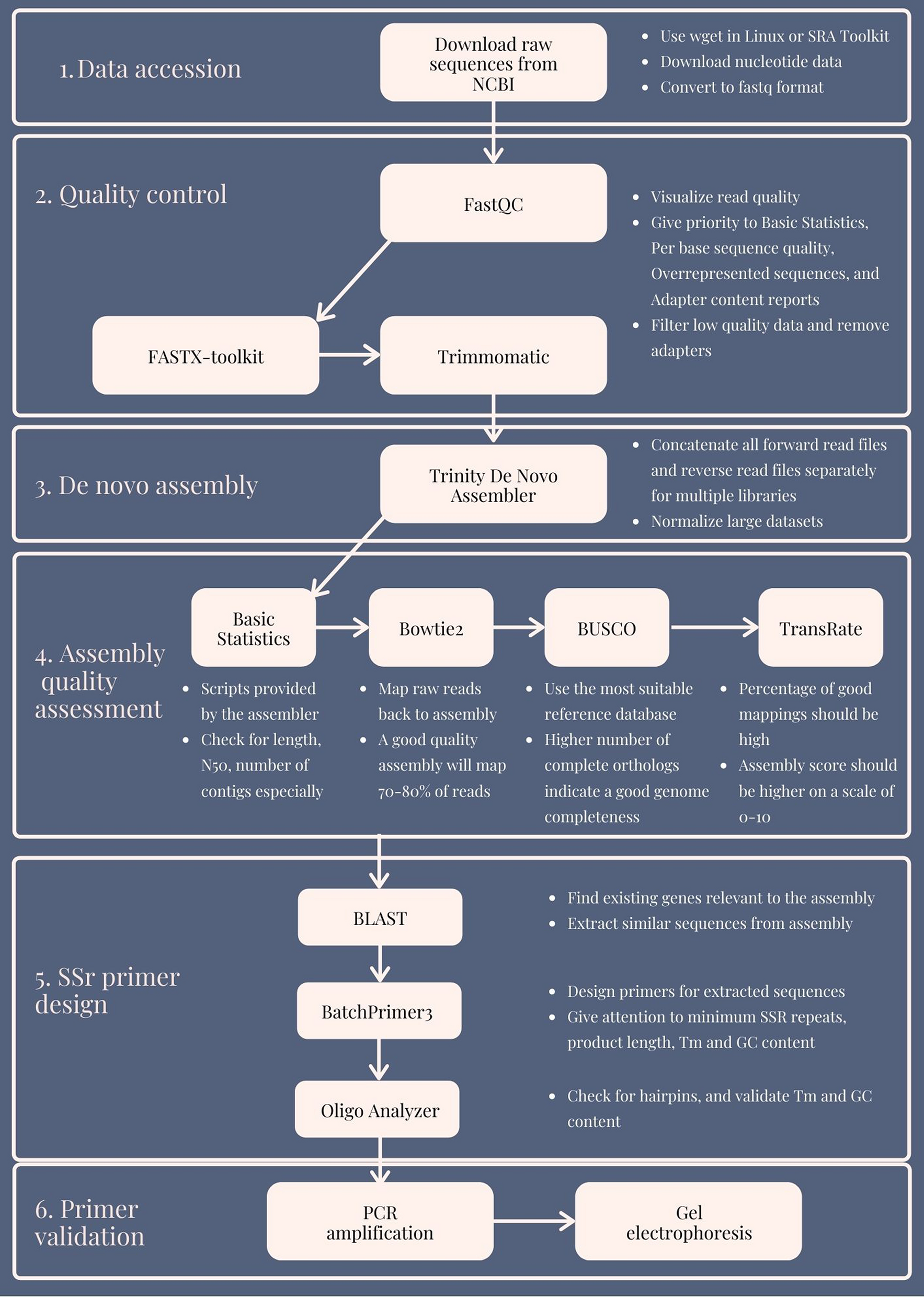
Raw transcriptomics data to gene specific SSRs: a validated free bioinformatics workflow for biologists | Scientific Reports

aTRAM 2.0: An Improved, Flexible Locus Assembler for NGS Data - Julie M Allen, Raphael LaFrance, Ryan A Folk, Kevin P Johnson, Robert P Guralnick, 2018

Review of General Algorithmic Features for Genome Assemblers for Next Generation Sequencers - ScienceDirect
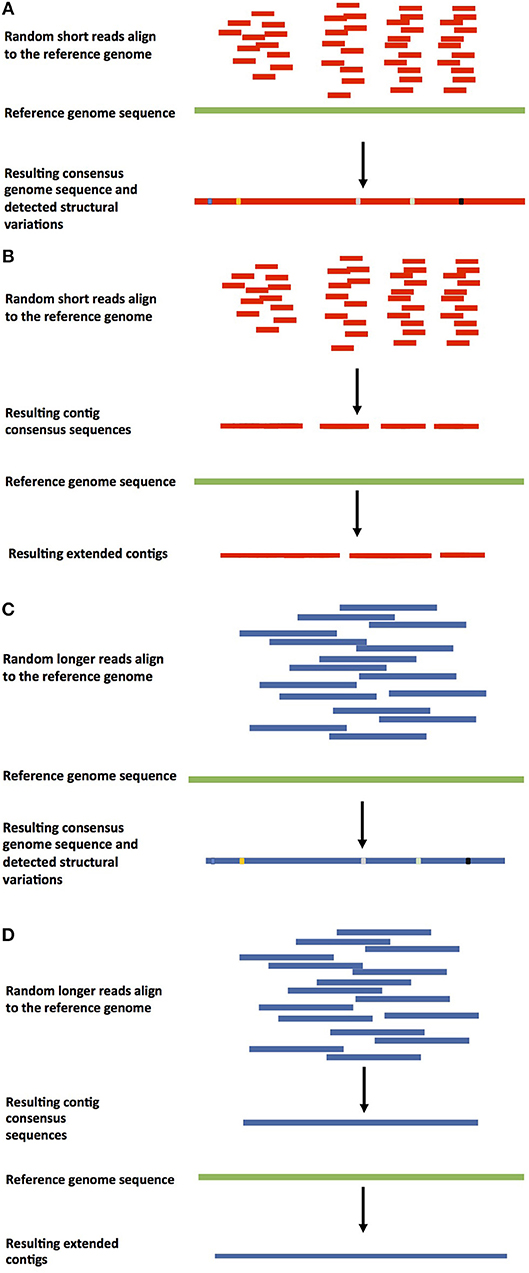
Next-Generation Sequence Assembly: Four Stages of Data Processing and Computational Challenges | PLOS Computational Biology
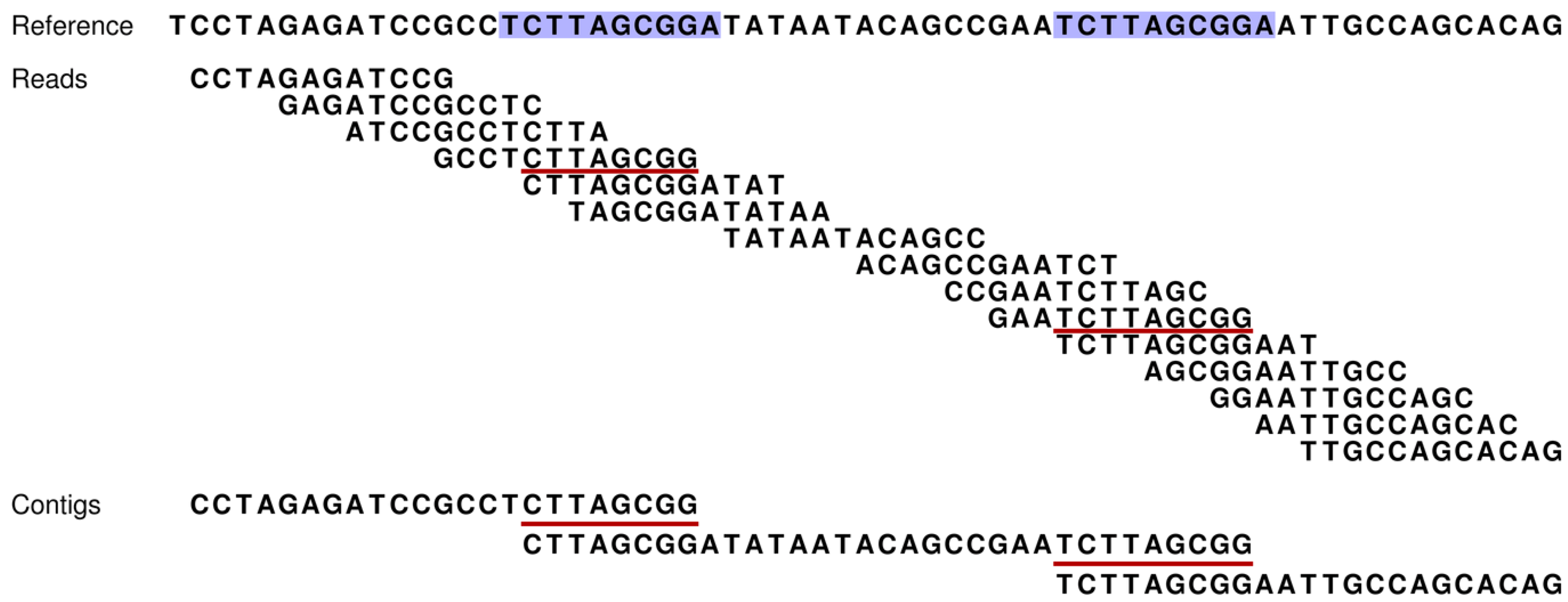
Genes | Free Full-Text | A Computer Simulator for Assessing Different Challenges and Strategies of de Novo Sequence Assembly

A field guide to whole‐genome sequencing, assembly and annotation - Ekblom - 2014 - Evolutionary Applications - Wiley Online Library
Next-Generation Sequence Assembly: Four Stages of Data Processing and Computational Challenges | PLOS Computational Biology

Review of General Algorithmic Features for Genome Assemblers for Next Generation Sequencers - ScienceDirect
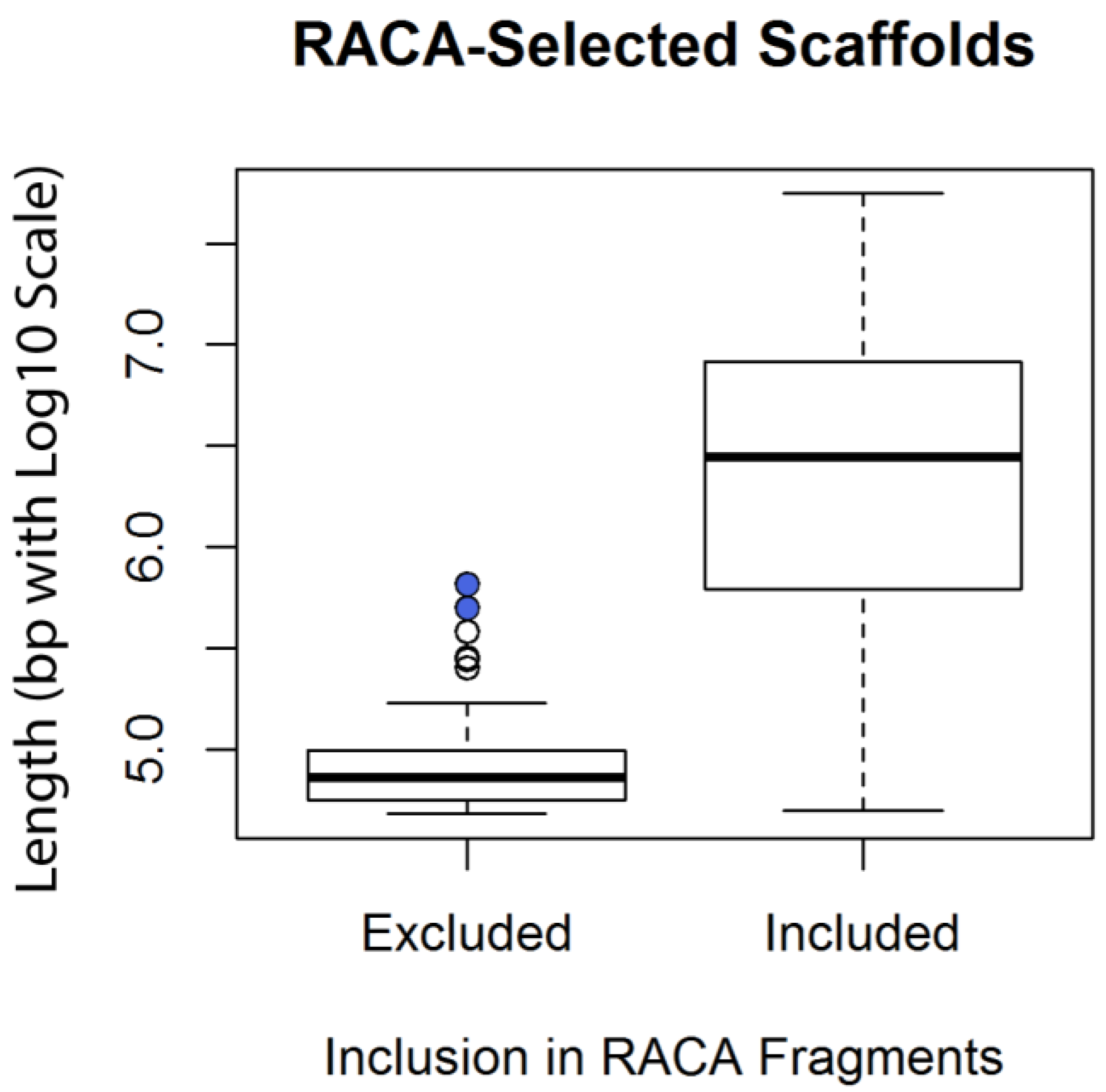
Genes | Free Full-Text | Construction of Red Fox Chromosomal Fragments from the Short-Read Genome Assembly
Next-Generation Sequence Assembly: Four Stages of Data Processing and Computational Challenges | PLOS Computational Biology
Optimizing Information in Next-Generation-Sequencing (NGS) Reads for Improving De Novo Genome Assembly | PLOS ONE
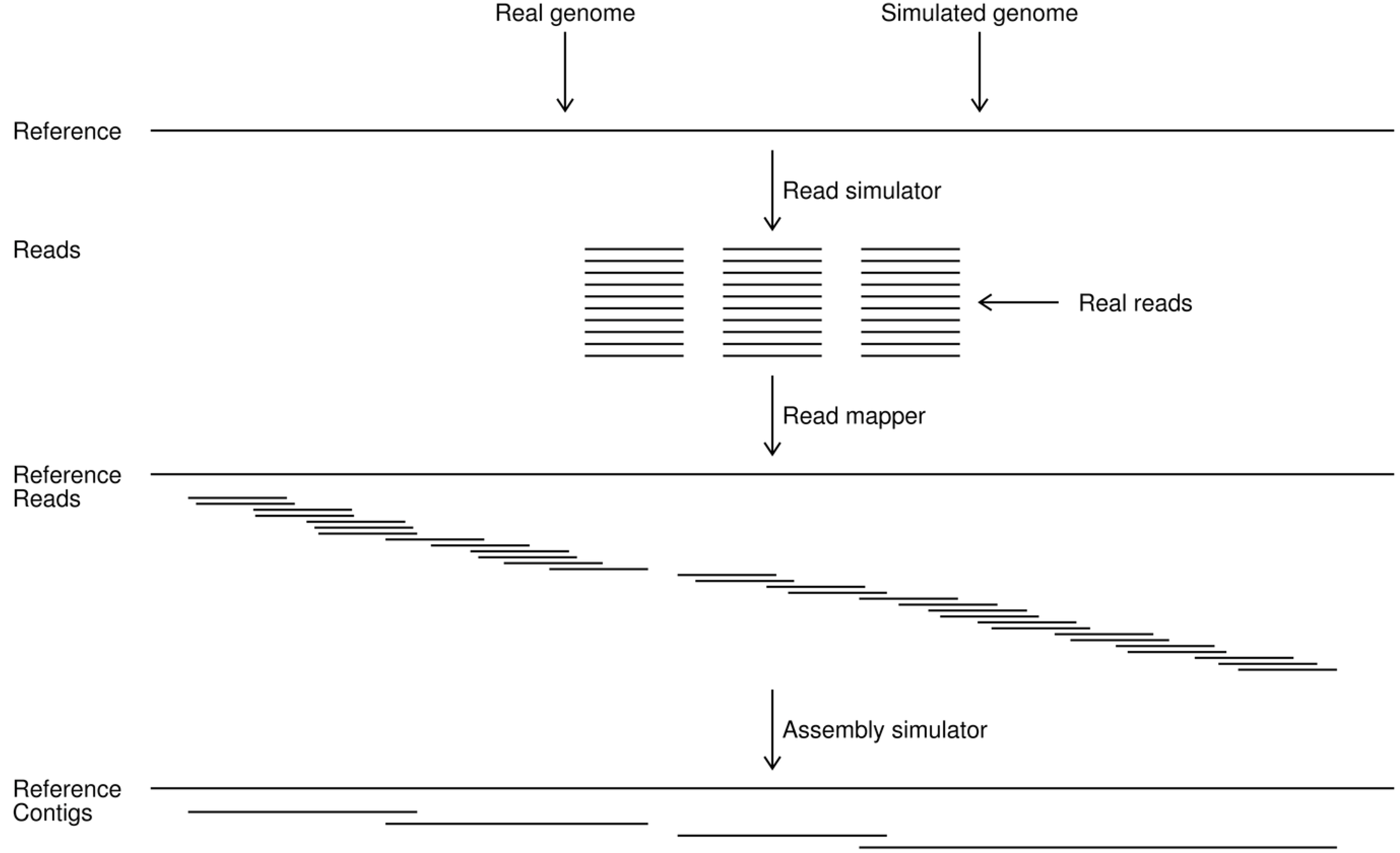


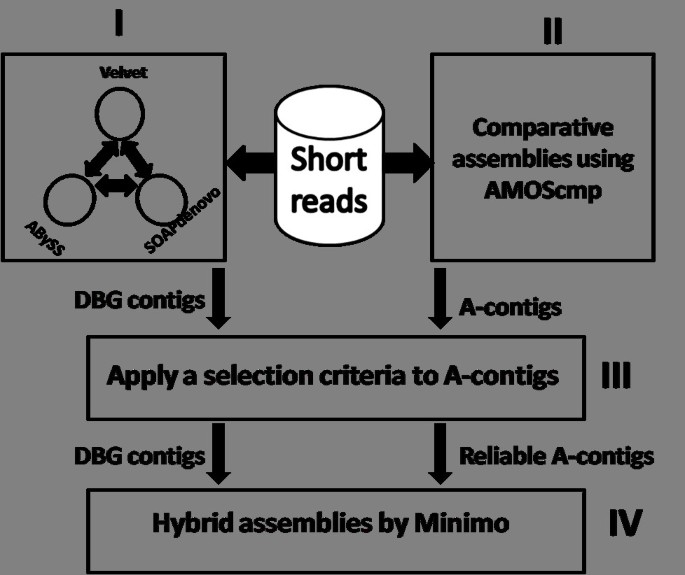

![PDF] Genome assembly reborn: recent computational challenges | Semantic Scholar PDF] Genome assembly reborn: recent computational challenges | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3fb111a7b2fd6725085ef184592b1d6b3308dba2/4-Figure1-1.png)